
python中的Mann-Kendall单调趋势检验--及原理说明_liucheng_zimozigreat的博客-CSDN博客_mann-kendall python
前提假设:
- 当没有趋势时,随时间获得的数据是独立同分布的。独立的假设是说数据随着时间不是连续相关的。
- 所获得的时间序列上的数据代表了采样时的真实条件。(样本具有代表性)
- 样本的采集、处理和测量方法提供了总体样本中的无偏且具有代表性的观测值。
pymannkendall的Python项目
什么是mann-kendall检验?
mann-kendall趋势检验(有时称为mk检验)用于分析时间序列数据的一致性增加或减少趋势(单调趋势)。这是一个非参数检验,这意味着它适用于所有分布(即数据不必满足正态性假设),但数据应该没有序列相关性。如果数据具有序列相关性,则可能在显著水平上影响(p值)。这可能会导致误解。为了克服这一问题,研究者提出了几种修正的mann-kendall检验(hamed和rao修正的mk检验、yue和wang修正的mk检验、预白化法修正的mk检验等)。季节性mann-kendall检验也被用来消除季节性的影响。
mann-kendall检验是一种强大的趋势检验,因此针对空间条件,发展了多元mk检验、区域mk检验、相关mk检验、部分mk检验等修正的mann-kendall检验。pymannkendal是非参数mann-kendall趋势分析的纯python实现,它集合了几乎所有类型的mann-kendall测试。目前,该软件包有11个mann-kendall检验和2个sen斜率估计函数。功能简介如下:
- 原始mann-kendall检验(原始_检验):原始mann-kendall检验是非参数检验,不考虑序列相关性或季节性影响。
- hamed和rao修正的mk检验(hamed和rao修正的mk检验):这个修正的mk检验由hamed和rao(1998)提出的解决序列自相关问题的方法。他们建议采用方差校正方法来改进趋势分析。用户可以通过在该函数中插入滞后数来考虑前n个显著滞后。默认情况下,它会考虑所有重要的延迟。
- Yue和Wang修正的MK检验(Yue-Wang_修正的检验):这也是Yue,S.,&Wang,C.Y.(2004)提出的考虑序列自相关的方差修正方法。用户还可以为计算设置所需的有效n滞后。
- 使用预白化方法的修正mk检验(预白化方法的修正):Yue和Wang(2002)建议在应用趋势检验之前使用预白化时间序列的检验。
- 使用无趋势预白化方法的修正mk试验(无趋势预白化试验):Yue和Wang(2002)也提出了在应用趋势试验之前去除趋势成分,然后对时间序列进行预白化的试验。
- 多变量mk检验(多变量检验):这是hirsch(1982)提出的多参数mk检验。他用这种方法进行季节性mk检验,把每个月作为一个参数。
- 季节性MK检验(季节性检验):对于季节性时间序列数据,Hirsch,R.M.,Slack,J.R.和Smith,R.A.(1982)提出了这个检验来计算季节性趋势。
- 区域mk检验(regional mk test):基于Hirsch(1982)提出的季节性mk检验,Helsel,D.R.和Frans,L.M.,(2006)建议采用区域mk检验来计算区域尺度的总体趋势。
- 相关多变量mk检验(相关多变量检验):hipel(1994)提出的参数相关的多变量mk检验。
- 相关季节性MK检验(相关季节性检验):当时间序列与前一个或多个月/季节显著相关时,使用Hipel(1994)提出的方法。
- 部分mk检验(部分_检验):在实际事件中,许多因素都会影响研究的主要响应参数,从而使趋势结果产生偏差。为了克服这个问题,libiseller(2002)提出了部分mk检验。它需要两个参数作为输入,一个是响应参数,另一个是独立参数。
- 泰尔-森斜率估计器(sen s-slope):泰尔(1950)和森(1968)提出的估计单调趋势幅度的方法。
- 季节sen斜率估计量(季节sen斜率):hipel(1994)提出的当数据具有季节性影响时估计单调趋势大小的方法。
功能详细信息:
所有mann-kendall检验函数的输入参数几乎相同。这些是:
- x:向量(列表、numpy数组或pandas系列)数据
- α:显著性水平(默认为0.05)
- 滞后:第一个有效滞后数(仅在hamed_rao_modification_test和yue_wang_modification_test中可用)
- 周期:季节性周期。月数据为12,周数据为52(仅在季节性测试中可用)
所有mann-kendall测试都返回一个命名元组,其中包含:
- 趋势:显示趋势(增加、减少或无趋势)
- h:真(如果趋势存在)或假(如果趋势不存在)
- p:显著性检验的p值
- z:标准化测试统计
- 陶:肯德尔陶
- s:Mann Kendal的分数
- 方差s:方差s
- 斜率:sen的斜率
sen的斜率函数需要数据向量。季节性sen的斜率也有可选的输入周期,默认值为12。两个sen的slope函数都只返回slope值。
Python pymannkendall包_程序模块 - PyPI - Python中文网
""" Created on 05 March 2018 Update on 26 July 2019 @author: Md. Manjurul Hussain Shourov version: 1.1 Approach: Vectorisation Citation: Hussain et al., (2019). pyMannKendall: a python package for non parametric Mann Kendall family of trend tests.. Journal of Open Source Software, 4(39), 1556, https://doi.org/10.21105/joss.01556 """ from __future__ import division import numpy as np from scipy.stats import norm, rankdata from collections import namedtuple # Supporting Functions # Data Preprocessing def __preprocessing(x): x = np.asarray(x) dim = x.ndim if dim == 1: c = 1 elif dim == 2: (n, c) = x.shape if c == 1: dim = 1 x = x.flatten() else: print('Please check your dataset.') return x, c # Missing Values Analysis def __missing_values_analysis(x, method = 'skip'): if method.lower() == 'skip': if x.ndim == 1: x = x[~np.isnan(x)] else: x = x[~np.isnan(x).any(axis=1)] n = len(x) return x, n # ACF Calculation def __acf(x, nlags): y = x - x.mean() n = len(x) d = n * np.ones(2 * n - 1) acov = (np.correlate(y, y, 'full') / d)[n - 1:] return acov[:nlags+1]/acov[0] # vectorization approach to calculate mk score, S def __mk_score(x, n): s = 0 demo = np.ones(n) for k in range(n-1): s = s + np.sum(demo[k+1:n][x[k+1:n] > x[k]]) - np.sum(demo[k+1:n][x[k+1:n] < x[k]]) return s # original Mann-Kendal's variance S calculation def __variance_s(x, n): # calculate the unique data unique_x = np.unique(x) g = len(unique_x) # calculate the var(s) if n == g: # there is no tie var_s = (n*(n-1)*(2*n+5))/18 else: # there are some ties in data tp = np.zeros(unique_x.shape) demo = np.ones(n) for i in range(g): tp[i] = np.sum(demo[x == unique_x[i]]) var_s = (n*(n-1)*(2*n+5) - np.sum(tp*(tp-1)*(2*tp+5)))/18 return var_s # standardized test statistic Z def __z_score(s, var_s): if s > 0: z = (s - 1)/np.sqrt(var_s) elif s == 0: z = 0 elif s < 0: z = (s + 1)/np.sqrt(var_s) return z # calculate the p_value def __p_value(z, alpha): # two tail test p = 2*(1-norm.cdf(abs(z))) h = abs(z) > norm.ppf(1-alpha/2) if (z < 0) and h: trend = 'decreasing' elif (z > 0) and h: trend = 'increasing' else: trend = 'no trend' return p, h, trend def __R(x): n = len(x) R = [] for j in range(n): i = np.arange(n) s = np.sum(np.sign(x[j] - x[i])) R.extend([(n + 1 + s)/2]) return np.asarray(R) def __K(x,z): n = len(x) K = 0 for i in range(n-1): j = np.arange(i,n) K = K + np.sum(np.sign((x[j] - x[i]) * (z[j] - z[i]))) return K # Original Sens Estimator def __sens_estimator(x): idx = 0 n = len(x) d = np.ones(int(n*(n-1)/2)) for i in range(n-1): j = np.arange(i+1,n) d[idx : idx + len(j)] = (x[j] - x[i]) / (j - i) idx = idx + len(j) return d def sens_slope(x): """ This method proposed by Theil (1950) and Sen (1968) to estimate the magnitude of the monotonic trend. Input: x: a one dimensional vector (list, numpy array or pandas series) data Output: slope: sen's slope Examples -------- >>> x = np.random.rand(120) >>> slope = sens_slope(x) """ x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') return np.median(__sens_estimator(x)) def seasonal_sens_slope(x, period=12): """ This method proposed by Hipel (1994) to estimate the magnitude of the monotonic trend, when data has seasonal effects. Input: x: a vector (list, numpy array or pandas series) data period: seasonal cycle. For monthly data it is 12, weekly data it is 52 (12 is the default) Output: slope: sen's slope Examples -------- >>> x = np.random.rand(120) >>> slope = seasonal_sens_slope(x, 12) """ x, c = __preprocessing(x) n = len(x) if x.ndim == 1: if np.mod(n,period) != 0: x = np.pad(x,(0,period - np.mod(n,period)), 'constant', constant_values=(np.nan,)) x = x.reshape(int(len(x)/period),period) x, n = __missing_values_analysis(x, method = 'skip') d = [] for i in range(period): d.extend(__sens_estimator(x[:,i])) return np.median(np.asarray(d)) def original_test(x, alpha = 0.05): """ This function checks the Mann-Kendall (MK) test (Mann 1945, Kendall 1975, Gilbert 1987). Input: x: a vector (list, numpy array or pandas series) data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.original_test(x,0.05) """ res = namedtuple('Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') s = __mk_score(x, n) var_s = __variance_s(x, n) Tau = s/(.5*n*(n-1)) z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) slope = sens_slope(x) return res(trend, h, p, z, Tau, s, var_s, slope) def hamed_rao_modification_test(x, alpha = 0.05, lag=None): """ This function checks the Modified Mann-Kendall (MK) test using Hamed and Rao (1998) method. Input: x: a vector (list, numpy array or pandas series) data alpha: significance level (0.05 default) lag: No. of First Significant Lags (default None, You can use 3 for considering first 3 lags, which also proposed by Hamed and Rao(1998)) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.hamed_rao_modification_test(x,0.05) """ res = namedtuple('Modified_Mann_Kendall_Test_Hamed_Rao_Approach', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') s = __mk_score(x, n) var_s = __variance_s(x, n) Tau = s/(.5*n*(n-1)) # Hamed and Rao (1998) variance correction if lag is None: lag = n else: lag = lag + 1 # detrending # x_detrend = x - np.multiply(range(1,n+1), np.median(x)) slope = sens_slope(x) x_detrend = x - np.arange(1,n+1) * slope I = rankdata(x_detrend) # account for autocorrelation acf_1 = __acf(I, nlags=lag-1) interval = norm.ppf(1 - alpha / 2) / np.sqrt(n) upper_bound = 0 + interval lower_bound = 0 - interval sni = 0 for i in range(1,lag): if (acf_1[i] <= upper_bound and acf_1[i] >= lower_bound): sni = sni else: sni += (n-i) * (n-i-1) * (n-i-2) * acf_1[i] n_ns = 1 + (2 / (n * (n-1) * (n-2))) * sni var_s = var_s * n_ns z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) return res(trend, h, p, z, Tau, s, var_s, slope) def yue_wang_modification_test(x, alpha = 0.05, lag=None): """ Input: This function checks the Modified Mann-Kendall (MK) test using Yue and Wang (2004) method. x: a vector (list, numpy array or pandas series) data alpha: significance level (0.05 default) lag: No. of First Significant Lags (default None, You can use 1 for considering first 1 lags, which also proposed by Yue and Wang (2004)) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.yue_wang_modification_test(x,0.05) """ res = namedtuple('Modified_Mann_Kendall_Test_Yue_Wang_Approach', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') s = __mk_score(x, n) var_s = __variance_s(x, n) Tau = s/(.5*n*(n-1)) # Yue and Wang (2004) variance correction if lag is None: lag = n else: lag = lag + 1 # detrending slope = sens_slope(x) x_detrend = x - np.arange(1,n+1) * slope # account for autocorrelation acf_1 = __acf(x_detrend, nlags=lag-1) idx = np.arange(1,lag) sni = np.sum((1 - idx/n) * acf_1[idx]) n_ns = 1 + 2 * sni var_s = var_s * n_ns z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) return res(trend, h, p, z, Tau, s, var_s, slope) def pre_whitening_modification_test(x, alpha = 0.05): """ This function checks the Modified Mann-Kendall (MK) test using Pre-Whitening method proposed by Yue and Wang (2002). Input: x: a vector (list, numpy array or pandas series) data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.pre_whitening_modification_test(x,0.05) """ res = namedtuple('Modified_Mann_Kendall_Test_PreWhitening_Approach', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') # PreWhitening acf_1 = __acf(x, nlags=1)[1] a = range(0, n-1) b = range(1, n) x = x[b] - x[a]*acf_1 n = len(x) s = __mk_score(x, n) var_s = __variance_s(x, n) Tau = s/(.5*n*(n-1)) z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) slope = sens_slope(x) return res(trend, h, p, z, Tau, s, var_s, slope) def trend_free_pre_whitening_modification_test(x, alpha = 0.05): """ This function checks the Modified Mann-Kendall (MK) test using the trend-free Pre-Whitening method proposed by Yue and Wang (2002). Input: x: a vector (list, numpy array or pandas series) data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.trend_free_pre_whitening_modification_test(x,0.05) """ res = namedtuple('Modified_Mann_Kendall_Test_Trend_Free_PreWhitening_Approach', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') # detrending slope = sens_slope(x) x_detrend = x - np.arange(1,n+1) * slope # PreWhitening acf_1 = __acf(x_detrend, nlags=1)[1] a = range(0, n-1) b = range(1, n) x = x_detrend[b] - x_detrend[a]*acf_1 n = len(x) x = x + np.arange(1,n+1) * slope s = __mk_score(x, n) var_s = __variance_s(x, n) Tau = s/(.5*n*(n-1)) z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) slope = sens_slope(x) return res(trend, h, p, z, Tau, s, var_s, slope) def multivariate_test(x, alpha = 0.05): """ This function checks the Multivariate Mann-Kendall (MK) test, which is originally proposed by R. M. Hirsch and J. R. Slack (1984) for the seasonal Mann-Kendall test. Later this method also used Helsel (2006) for Regional Mann-Kendall test. Input: x: a matrix of data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.multivariate_test(x,0.05) """ res = namedtuple('Multivariate_Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) s = 0 var_s = 0 denom = 0 x, c = __preprocessing(x) # x, n = __missing_values_analysis(x, method = 'skip') # It makes all column at the same size for i in range(c): if c == 1: x_new, n = __missing_values_analysis(x, method = 'skip') # It makes all column at deferent size else: x_new, n = __missing_values_analysis(x[:,i], method = 'skip') # It makes all column at deferent size s = s + __mk_score(x_new, n) var_s = var_s + __variance_s(x_new, n) denom = denom + (.5*n*(n-1)) Tau = s/denom z = __z_score(s, var_s) p, h, trend = __p_value(z, alpha) slope = seasonal_sens_slope(x, period = c) return res(trend, h, p, z, Tau, s, var_s, slope) def seasonal_test(x, period = 12, alpha = 0.05): """ This function checks the Seasonal Mann-Kendall (MK) test (Hirsch, R. M., Slack, J. R. 1984). Input: x: a vector of data period: seasonal cycle. For monthly data it is 12, weekly data it is 52 (12 is the default) alpha: significance level (0.05 is the default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.seasonal_test(x,0.05) """ res = namedtuple('Seasonal_Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) n = len(x) if x.ndim == 1: if np.mod(n,period) != 0: x = np.pad(x,(0,period - np.mod(n,period)), 'constant', constant_values=(np.nan,)) x = x.reshape(int(len(x)/period),period) trend, h, p, z, Tau, s, var_s, slope = multivariate_test(x, alpha = alpha) return res(trend, h, p, z, Tau, s, var_s, slope) def regional_test(x, alpha = 0.05): """ This function checks the Regional Mann-Kendall (MK) test (Helsel 2006). Input: x: a matrix of data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.regional_test(x,0.05) """ res = namedtuple('Regional_Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) trend, h, p, z, Tau, s, var_s, slope = multivariate_test(x) return res(trend, h, p, z, Tau, s, var_s, slope) def correlated_multivariate_test(x, alpha = 0.05): """ This function checks the Correlated Multivariate Mann-Kendall (MK) test (Libiseller and Grimvall (2002)). Input: x: a matrix of data alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.correlated_multivariate_test(x,0.05) """ res = namedtuple('Correlated_Multivariate_Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) x, n = __missing_values_analysis(x, method = 'skip') s = 0 denom = 0 for i in range(c): s = s + __mk_score(x[:,i], n) denom = denom + (.5*n*(n-1)) Tau = s/denom Gamma = np.ones([c,c]) for i in range(1,c): for j in range(i): k = __K(x[:,i], x[:,j]) ri = __R(x[:,i]) rj = __R(x[:,j]) Gamma[i,j] = (k + 4 * np.sum(ri * rj) - n*(n+1)2)/3 Gamma[j,i] = Gamma[i,j] for i in range(c): k = __K(x[:,i], x[:,i]) ri = __R(x[:,i]) rj = __R(x[:,i]) Gamma[i,i] = (k + 4 * np.sum(ri * rj) - n*(n+1)2)/3 var_s = np.sum(Gamma) z = s / np.sqrt(var_s) p, h, trend = __p_value(z, alpha) slope = seasonal_sens_slope(x, period=c) return res(trend, h, p, z, Tau, s, var_s, slope) def correlated_seasonal_test(x, period = 12 ,alpha = 0.05): """ This function checks the Correlated Seasonal Mann-Kendall (MK) test (Hipel [1994] ). Input: x: a matrix of data period: seasonal cycle. For monthly data it is 12, weekly data it is 52 (12 is default) alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.correlated_seasonal_test(x,0.05) """ res = namedtuple('Correlated_Seasonal_Mann_Kendall_test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x, c = __preprocessing(x) n = len(x) if x.ndim == 1: if np.mod(n,period) != 0: x = np.pad(x,(0,period - np.mod(n,period)), 'constant', constant_values=(np.nan,)) x = x.reshape(int(len(x)/period),period) trend, h, p, z, Tau, s, var_s, slope = correlated_multivariate_test(x) return res(trend, h, p, z, Tau, s, var_s, slope) def partial_test(x, alpha = 0.05): """ This function checks the Partial Mann-Kendall (MK) test (Libiseller and Grimvall (2002)). Input: x: a matrix with 2 columns alpha: significance level (0.05 default) Output: trend: tells the trend (increasing, decreasing or no trend) h: True (if trend is present) or False (if trend is absence) p: p-value of the significance test z: normalized test statistics Tau: Kendall Tau s: Mann-Kendal's score var_s: Variance S slope: sen's slope Examples -------- >>> import pymannkendall as mk >>> x = np.random.rand(1000) >>> trend,h,p,z,tau,s,var_s,slope = mk.partial_test(x,0.05) """ res = namedtuple('Partial_Mann_Kendall_Test', ['trend', 'h', 'p', 'z', 'Tau', 's', 'var_s', 'slope']) x_old, c = __preprocessing(x) x_old, n = __missing_values_analysis(x_old, method = 'skip') if c != 2: raise ValueError('Partial Mann Kendall test required two parameters/columns. Here column no ' + str(c) + ' is not equal to 2.') x = x_old[:,0] y = x_old[:,1] x_score = __mk_score(x, n) y_score = __mk_score(y, n) k = __K(x, y) rx = __R(x) ry = __R(y) sigma = (k + 4 * np.sum(rx * ry) - n*(n+1)2)/3 rho = sigma / (n*(n-1)*(2*n+5)/18) s = x_score - rho * y_score var_s = (1 - rho2) * (n*(n-1)*(2*n+5))/18 Tau = x_score/(.5*n*(n-1)) z = s / np.sqrt(var_s) p, h, trend = __p_value(z, alpha) slope = sens_slope(x) return res(trend, h, p, z, Tau, s, var_s, slope)讯享网
批量逐点检验(未处理自相关)
scipy.stats.kendalltau() 函数
Kendall's tau-b(肯德尔)等级相关系数:用于反映分类变量相关性的指标,适用于两个分类变量(时间—水文要素)均为有序分类的情况。对相关的有序变量进行非参数相关检验;取值范围在-1-1之间,此检验适合于正方形表格;


scipy.stats.kendalltau — SciPy v0.19.1 Reference Guide
讯享网from scipy import stats import pandas as pd import numpy as np data = pd.read_csv(r"C:\Users\Leon\Desktop\Pre.csv") #print (data) 38行*994列(38年994个cell) x = range(38) print (x) y = np.zeros((0)) for j in range(994): b = stats.kendalltau(x,data.values[:,j]) MK检验,结果包含两个参数:tau, p_value y = np.append(y, b, axis=0) print(b) print(type(y)) #np.savetxt("C:/Users/Leon/Desktop/P.txt",y) 保存ndarray类型数据
稳健回归(Robustness regression)
最小二乘法的弊端
比如下图中所示,数据中存在一个异常点,如果不剔除改点,适用OLS方法来做回归的话,那么就会得到途中红色的那条线;如果将这个异常点剔除掉的话,那么就可以得到图中蓝色的那条线。显然,蓝色的线比红色的线对数据有更强的解释性,这就是OLS在做回归分析时候的弊端。
当然,可以考虑在做回归分析之前,对数据做预处理,剔除掉那些异常点。但是,在实际的数据中,存在两个问题:
异常点并不能很好的确定,并没有一个很好的标准用于确定哪些点是异常点
即便确定了异常点,但这些被确定为异常的点,真的是错误的数据吗?很有可能这看似异常的点,就是原始模型的数据,如果是这样的话,那么这些异常的点就会带有大量的原始模型的信息,剔除之后就会丢失大量的信息。
再比如下面这幅图,其中红色的都是异常点,但是很难从数据中剔除出去。
稳健回归
Breakdown point
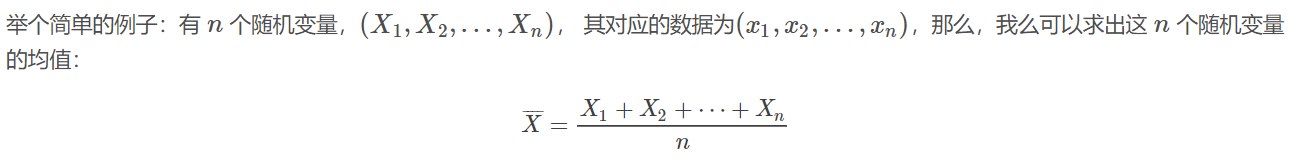
这个均值估计器的Breakdown point 为0,因为使任意一个xixi变成足够大的脏数据之后,上面估计出来的均值,就不再正确了。
毫无疑问,Breakdown point越大,估计器就越稳健。
Breakdown point 是不可能达到 50% 的,因为如果总体样本中超过一半的数据是脏数据了,那么从统计上来说,就无法将样本中的隐藏分布和脏数据的分布给区分开来。
本文主要介绍两种稳健回归模型:RANSAC(RANdom SAmple Consensus 随机采样一致性)和Theil-Sen estimator。
RANSAC随机采样一致性算法

RANSAC算法是从输入样本集合的内点的随机子集中学习模型。
RANSAC算法是一个非确定性算法(non-deterministic algorithm),这个算法只能得以一定的概率得到一个还不错的结果,在基本模型已定的情况下,结果的好坏程度主要取决于算法最大的迭代次数。
RANSAC算法在线性和非线性回归中都得到了广泛的应用,而其最典型也是最成功的应用,莫过于在图像处理中处理图像拼接问题,这部分在Opencv中有相关的实现。
从总体上来讲,RANSAC算法将输入样本分成了两个大的子集:内点(inliers)和外点(outliers)。其中内点的数据分布会受到噪声的影响;而外点主要来自于错误的测量手段或者是对数据错误的假设。而RANSAC算法最终的结果是基于算法所确定的内点集合得到的。
# -*- coding: utf-8 -*- """ author : time : 2016-07-07-15-36 """ import numpy as np import time from sklearn import linear_model,datasets import matplotlib.pyplot as plt # 产生数据样本点集合 # 样本点的特征X维度为1维,输出y的维度也为1维 # 输出是在输入的基础上加入了高斯噪声N(0,10) # 产生的样本点数目为1000个 n_samples = 1000 X, y, coef = datasets.make_regression(n_samples=n_samples, n_features=1, n_informative=1, noise=10, coef=True, random_state=0) # 将上面产生的样本点中的前50个设为异常点(外点) # 即:让前50个点偏离原来的位置,模拟错误的测量带来的误差 n_outliers = 50 np.random.seed(int(time.time()) % 100) X[:n_outliers] = 3 + 0.5 * np.random.normal(size=(n_outliers, 1)) y[:n_outliers] = -3 + 0.5 * np.random.normal(size=n_outliers) # 用普通线性模型拟合X,y model = linear_model.LinearRegression() model.fit(X, y) # 使用RANSAC算法拟合X,y model_ransac = linear_model.RANSACRegressor(linear_model.LinearRegression()) model_ransac.fit(X, y) inlier_mask = model_ransac.inlier_mask_ outlier_mask = np.logical_not(inlier_mask) # 使用一般回归模型和RANSAC算法分别对测试数据做预测 line_X = np.arange(-5, 5) line_y = model.predict(line_X[:, np.newaxis]) line_y_ransac = model_ransac.predict(line_X[:, np.newaxis]) print "真实数据参数:", coef print "线性回归模型参数:", model.coef_ print "RANSAC算法参数: ", model_ransac.estimator_.coef_ plt.plot(X[inlier_mask], y[inlier_mask], '.g', label='Inliers') plt.plot(X[outlier_mask], y[outlier_mask], '.r', label='Outliers') plt.plot(line_X, line_y, '-k', label='Linear Regression') plt.plot(line_X, line_y_ransac, '-b', label="RANSAC Regression") plt.legend(loc='upper left') plt.show()
运行结果为:
讯享网真实数据参数: 82. 线性回归模型参数: [ 55.] RANSAC算法参数: [ 82.0]
Theil-Sen Regression 泰尔森回归
Theil-Sen回归是一个参数中值估计器,它适用泛化中值,对多维数据进行估计,因此其对多维的异常点(outliers 外点)有很强的稳健性。

在实践中发现,随着数据特征维度的提升,Theil-Sen回归的效果不断的下降,在高维数据中,Theil-Sen回归的效果有时甚至还不如OLS(最小二乘)。
在之间的文章《线性回归》中讨论过,OLS方法是渐进无偏的,Theil-Sen方法在渐进无偏方面和OLS性能相似。和OLS方法不同的是,Theil-Sen方法是一种非参数方法,其对数据的潜在分布不做任何的假设。Theil-Sen方法是一种基于中值的估计其,所以其对异常点有更强的稳健性。
在单变量回归问题中,Theil-Sen方法的Breakdown point为29.3%,也就是说,Theil-Sen方法可以容忍29.3%的数据是outliers。
# -*- coding: utf-8 -*- """ @author : @time ;2016-07-08_08-50 Theil-Sen 回归 本例生成一个数据集,然后在该数据集上测试Theil-Sen回归 """ print __doc__ import time import numpy as np import matplotlib.pyplot as plt from sklearn.linear_model import LinearRegression, TheilSenRegressor,\ RANSACRegressor estimators = [('OLS', LinearRegression()), ('Theil-Sen', TheilSenRegressor())] # 异常值仅仅出现在y轴 np.random.seed((int)(time.time() % 100)) n_samples = 200 # 线性模型的函数形式为: y = 3 * x + N(2, .1 2) x = np.random.randn(n_samples) w = 3. c = 2. noise = c + 0.1 * np.random.randn(n_samples) y = w * x + noise # 加入10%的异常值,最后20个值称为异常值 y[-20:] += -20 * x[-20:] X = x[:, np.newaxis] plt.plot(X, y, 'k+', mew=2, ms=8) line_x = np.array([-3, 3]) for name, estimator in estimators: t0 = time.time() estimator.fit(X, y) elapsed_time = time.time() - t0 y_pred = estimator.predict(line_x.reshape(2, 1)) plt.plot(line_x, y_pred, label='%s (fit time: %.2fs)' %(name, elapsed_time)) plt.axis('tight') plt.legend(loc='upper left') plt.show() 


版权声明:本文内容由互联网用户自发贡献,该文观点仅代表作者本人。本站仅提供信息存储空间服务,不拥有所有权,不承担相关法律责任。如发现本站有涉嫌侵权/违法违规的内容,请联系我们,一经查实,本站将立刻删除。
如需转载请保留出处:https://51itzy.com/kjqy/49265.html