首先我们要定义所有样本的分型我们从TCGAbiolinks 中获取
library(TCGAbiolinks) subtypes <- PanCancerAtlas_subtypes() DT::datatable(subtypes, filter = 'top', options = list(scrollX = TRUE, keys = TRUE, pageLength = 5), rownames = FALSE) write.table(subtypes,file="sub.txt",quote=F,sep="\t") 讯享网
这里获取的是TCGA所有样本的分型,需要再次筛选你需要的分型 ,这里以BRCA为例

讯享网
这里的文档要整理成这个样子
2 下面一步是为了进行数据的清洗
根据给的TCGA的样本我们进行划分,区分的97个case是有对照
首先清洗正常组的
讯享网rt <- read.table('2.txt',sep = '\t',header = T,check.names = F) 导入的是上面表格的数据 rownames(rt) <- rt[,1] tcga_data <- read.table('3.txt',sep = '\t',header = T,check.names = F)导入99对那个数据 rownames(tcga_data) <- tcga_data[,1] tcga_data <- tcga_data[,-1] group_list=ifelse(as.numeric(substr(colnames(tcga_data),14,15)) < 10,'tumor','normal')定义TCGA分组 tcga_normal <- tcga_data[, group_list == 'normal']筛选正常组的数据 rt1=rt[match(colnames(tcga_normal),rt$pan.samplesID),] 查看ID是否在样本分型的数据集中 rt2=rt1[rt1$Subtype_mRNA%in%c("Normal"),]提取分型为正常的样本ID tcga_normal_need=tcga_normal[,colnames(tcga_normal)%in%rt2$pan.samplesID]整理样本分型为正常的样本 > dim(tcga_normal_need) [1] 60483 8797个中87的的癌旁是normal
2 接下来根据癌旁的结果选择对应癌做分析
首先我们看一下对应的癌是什么分型

tcga_tumor<- tcga_data[, group_list == 'tumor'] tcga_tumor_need <- tcga_tumor[,((substr(colnames(tcga_tumor),1,12))%in%(substr(rownames(rt2),1,12)))]根据正常样本TCGA前12位来提取相同的向量的肿瘤的样本 tcga_need <- cbind(tcga_normal_need,tcga_tumor_need)合并正常的与肿瘤 write.table(tcga_need,file = 'tcga_need.txt',sep = '\t',quote = F) dim(tcga_need) > dim(tcga_need) [1] 60483 174 3 我们对肿瘤样本的分型进行定义
讯享网rt3=rt[match(colnames(tcga_tumor_need),rt$pan.samplesID),]匹配ID并得到所有ID分型 LumA1 <-rt3[rt3$Subtype_mRNA%in%c("LumA"),]LumA分型 LumB1 <- rt3[rt3$Subtype_mRNA%in%c("LumB"),] Her21<- rt3[rt3$Subtype_mRNA%in%c("Her2"),] Bascal1<- rt3[rt3$Subtype_mRNA%in%c("Basal"),] Normal1<- rt3[rt3$Subtype_mRNA%in%c("Normal"),] LumA=tcga_need[,colnames(tcga_need)%in%LumA1$pan.samplesID ]LumA分型的数据 LumB=tcga_need[,colnames(tcga_need)%in%LumB1$pan.samplesID ] Her2=tcga_need[,colnames(tcga_need)%in%Her21$pan.samplesID ] Normal=tcga_need[,colnames(tcga_need)%in%Normal1$pan.samplesID ] Bascal=tcga_need[,colnames(tcga_need)%in%Bascal1$pan.samplesID ] data=cbind(LumA,LumB,Her2,Bascal,Normal)按照顺序进行分型的合并
这里展示一次分型对于肿瘤样本的分型接下来做PCA
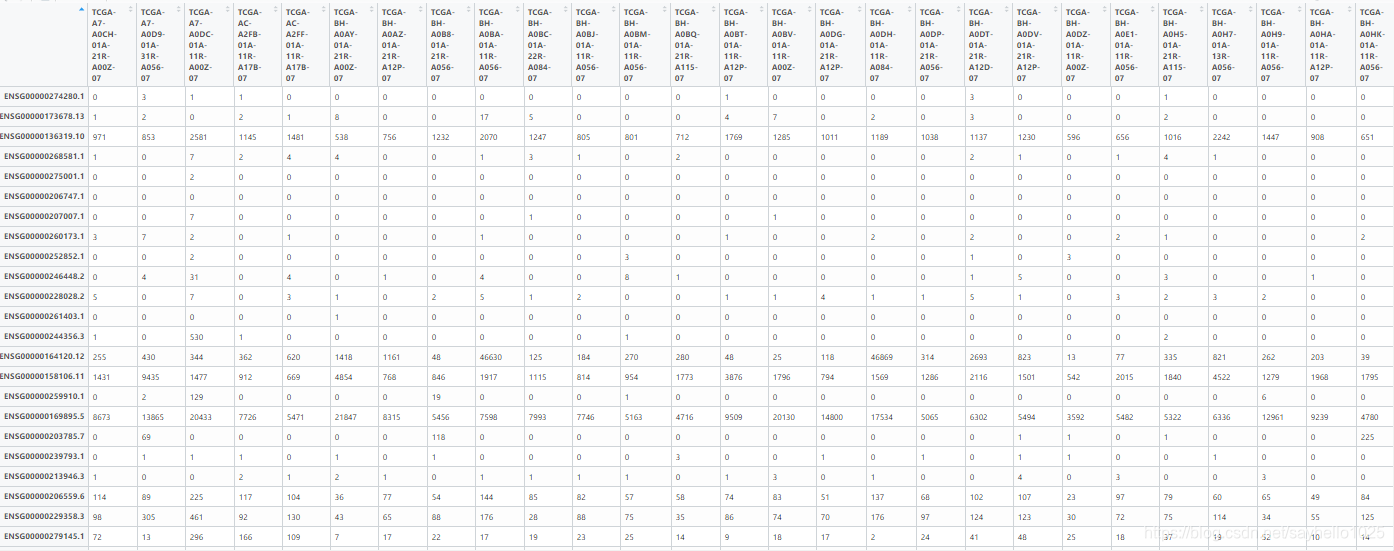
这是下面输入的DATA
library("FactoMineR") library("factoextra") library(tidyverse) setwd("") #设置工作目录 data <- read.table("sample.txt", header=T, row.names=NULL,sep="\t")数据导入 rownames_data <- make.names(data[,1],unique=T)下面机组都是对数据的整理如果是接着做就不用了 data <- data[,-1,drop=F] rownames(data) <- rownames_data data <- data[rowSums(data) =!0,]如果是上面图片的形式你可以从这一步开始,行不等于0 data <- data[apply(data, 1, var)!=0,] 列的方差不为0 pca <- prcomp(t(data),scale = TRUE)pca的核心 var_explained <- pca$sdev^2/sum(pca$sdev^2) write.table(pca$rotation,file="PC.xls",quote=F,sep="\t") #输出特征向量 write.table(predict(pca),file="newTab.xls",quote=F,sep="\t") #输出新表 pca.sum=summary(pca) write.table(pca.sum$importance,file="importance.xls",quote=F,sep="\t")#输出PC比重 pca$x %>% as.data.frame %>% ggplot(aes(x=PC1,y=PC2)) + geom_point(size=4) + theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%")) + theme(legend.position="top") pca$x %>% as.data.frame %>% rownames_to_column("continent_letter") %>% separate(continent_letter,c("continent")) %>% ggplot(aes(x=PC1,y=PC2)) + geom_point(aes(color=continent),size=4) + theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%")) + theme(legend.position="top") pca$x %>% as.data.frame %>% rownames_to_column("continent_typle") %>% separate(continent_typle,c("continent", "typle")) %>% ggplot(aes(x=PC1,y=PC2, label=typle, color=continent)) + geom_label(aes(fill = continent), colour = "white", fontface = "bold")+ theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%"))+ theme(legend.position="top") pdf(file="pcaBarplot.pdf",width=15) #柱状图 barplot(pca.sum$importance[2,]*100,xlab="PC",ylab="percent",col="skyblue") dev.off() pdf(file="pcaPlot.pdf",width=15) #碎石图 plot(pca.sum$importance[2,]*100,type="o",col="red",xlab="PC",ylab="percent") dev.off() library(ggplot2) 下面的分组是基于你前面data数据的顺序 group=c(rep("LumA",44),rep("LumB",17),rep("Her2",9),rep("Basal",14),rep("Normal",2)) #需要大家输入 pcaPredict=predict(pca) PCA = data.frame(PCA1 = pcaPredict[,1], PCA2 = pcaPredict[,2],group=group) PCA.mean=aggregate(PCA[,1:2],list(group=PCA$group),mean) #定义椭圆 veganCovEllipse<-function (cov, center = c(0, 0), scale = 1, npoints = 100) { theta <- (0:npoints) * 2 * pi/npoints Circle <- cbind(cos(theta), sin(theta)) t(center + scale * t(Circle %*% chol(cov))) } df_ell <- data.frame() for(g in levels(PCA$group)){ df_ell <- rbind(df_ell, cbind(as.data.frame(with(PCA[PCA$group==g,], veganCovEllipse(cov.wt(cbind(PCA1,PCA2), wt=rep(1/length(PCA1),length(PCA1)))$cov, center=c(mean(PCA1),mean(PCA2))))),group=g)) } pdf(file="PCA2d.pdf") ggplot(data = PCA, aes(PCA1, PCA2)) + geom_point(aes(color = group)) + geom_path(data=df_ell, aes(x=PCA1, y=PCA2,colour=group), size=1, linetype=2)+ annotate("text",x=PCA.mean$PCA1,y=PCA.mean$PCA2,label=PCA.mean$group)+ theme_bw()+ theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank()) dev.off() 讯享网library(TCGAbiolinks) subtypes <- PanCancerAtlas_subtypes() DT::datatable(subtypes, filter = 'top', options = list(scrollX = TRUE, keys = TRUE, pageLength = 5), rownames = FALSE) write.table(subtypes,file="sub.txt",quote=F,sep="\t") rt <- read.table('2.txt',sep = '\t',header = T,check.names = F) rownames(rt) <- rt[,1] tcga_data <- read.table('3.txt',sep = '\t',header = T,check.names = F) rownames(tcga_data) <- tcga_data[,1] tcga_data <- tcga_data[,-1] group_list=ifelse(as.numeric(substr(colnames(tcga_data),14,15)) < 10,'tumor','normal') tcga_normal <- tcga_data[, group_list == 'normal'] rt1=rt[match(colnames(tcga_normal),rt$pan.samplesID),] rt2=rt1[rt1$Subtype_mRNA%in%c("Normal"),] tcga_normal_need=tcga_normal[,colnames(tcga_normal)%in%rt2$pan.samplesID] dim(tcga_normal_need) tcga_tumor<- tcga_data[, group_list == 'tumor'] tcga_tumor_need <- tcga_tumor[,((substr(colnames(tcga_tumor),1,12))%in%(substr(rownames(rt2),1,12)))] tcga_need <- cbind(tcga_normal_need,tcga_tumor_need) write.table(tcga_need,file = 'tcga_need.txt',sep = '\t',quote = F) dim(tcga_need) rt3=rt[match(colnames(tcga_tumor_need),rt$pan.samplesID),] LumA1 <-rt3[rt3$Subtype_mRNA%in%c("LumA"),] LumB1 <- rt3[rt3$Subtype_mRNA%in%c("LumB"),] Her21<- rt3[rt3$Subtype_mRNA%in%c("Her2"),] Bascal1<- rt3[rt3$Subtype_mRNA%in%c("Basal"),] Normal1<- rt3[rt3$Subtype_mRNA%in%c("Normal"),] LumA=tcga_need[,colnames(tcga_need)%in%LumA1$pan.samplesID ] LumB=tcga_need[,colnames(tcga_need)%in%LumB1$pan.samplesID ] Her2=tcga_need[,colnames(tcga_need)%in%Her21$pan.samplesID ] Normal=tcga_need[,colnames(tcga_need)%in%Normal1$pan.samplesID ] Bascal=tcga_need[,colnames(tcga_need)%in%Bascal1$pan.samplesID ] data=cbind(LumA,LumB,Her2,Bascal,Normal) data <- data[rowSums(data)>0,] data <- data[apply(data, 1, var)!=0,] pca <- prcomp(t(data),scale = TRUE) var_explained <- pca$sdev^2/sum(pca$sdev^2) write.table(pca$rotation,file="PC.xls",quote=F,sep="\t") #输出特征向量 write.table(predict(pca),file="newTab.xls",quote=F,sep="\t") #输出新表 pca.sum=summary(pca) write.table(pca.sum$importance,file="importance.xls",quote=F,sep="\t")#输出PC比重 pca$x %>% as.data.frame %>% ggplot(aes(x=PC1,y=PC2)) + geom_point(size=4) + theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%")) + theme(legend.position="top") pca$x %>% as.data.frame %>% rownames_to_column("continent_letter") %>% separate(continent_letter,c("continent")) %>% ggplot(aes(x=PC1,y=PC2)) + geom_point(aes(color=continent),size=4) + theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%")) + theme(legend.position="top") pca$x %>% as.data.frame %>% rownames_to_column("continent_typle") %>% separate(continent_typle,c("continent", "typle")) %>% ggplot(aes(x=PC1,y=PC2, label=typle, color=continent)) + geom_label(aes(fill = continent), colour = "white", fontface = "bold")+ theme_bw(base_size=32) + labs(x=paste0("PC1: ",round(var_explained[1]*100,1),"%"), y=paste0("PC2: ",round(var_explained[2]*100,1),"%"))+ theme(legend.position="top") pdf(file="pcaBarplot.pdf",width=15) #柱状图 barplot(pca.sum$importance[2,]*100,xlab="PC",ylab="percent",col="skyblue") dev.off() pdf(file="pcaPlot.pdf",width=15) #碎石图 plot(pca.sum$importance[2,]*100,type="o",col="red",xlab="PC",ylab="percent") dev.off() library(ggplot2) group=c(rep("LumA",44),rep("LumB",17),rep("Her2",9),rep("Basal",14),rep("Normal",2)) pcaPredict=predict(pca) PCA = data.frame(PCA1 = pcaPredict[,1], PCA2 = pcaPredict[,2],group=group) PCA.mean=aggregate(PCA[,1:2],list(group=PCA$group),mean) #定义椭圆 veganCovEllipse<-function (cov, center = c(0, 0), scale = 1, npoints = 100) { theta <- (0:npoints) * 2 * pi/npoints Circle <- cbind(cos(theta), sin(theta)) t(center + scale * t(Circle %*% chol(cov))) } df_ell <- data.frame() for(g in levels(PCA$group)){ df_ell <- rbind(df_ell, cbind(as.data.frame(with(PCA[PCA$group==g,], veganCovEllipse(cov.wt(cbind(PCA1,PCA2), wt=rep(1/length(PCA1),length(PCA1)))$cov, center=c(mean(PCA1),mean(PCA2))))),group=g))} pdf(file="PCA2d.pdf") ggplot(data = PCA, aes(PCA1, PCA2)) + geom_point(aes(color = group)) + geom_path(data=df_ell, aes(x=PCA1, y=PCA2,colour=group), size=1, linetype=)+ annotate("text",x=PCA.mean$PCA1,y=PCA.mean$PCA2,label=PCA.mean$group)+ theme_bw()+ theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank()) dev.off() pdf(file="PCA2d.pdf") ggplot(data = PCA, aes(PCA1, PCA2)) + geom_point(aes(color = group)) + geom_path(data=df_ell, aes(x=PCA1, y=PCA2,colour=group), size=1, linetype=2)+ annotate("text",x=PCA.mean$PCA1,y=PCA.mean$PCA2,label=PCA.mean$group)+ theme_bw()+ theme(panel.grid.major = element_blank(), panel.grid.minor = element_blank()) dev.off()

版权声明:本文内容由互联网用户自发贡献,该文观点仅代表作者本人。本站仅提供信息存储空间服务,不拥有所有权,不承担相关法律责任。如发现本站有涉嫌侵权/违法违规的内容,请联系我们,一经查实,本站将立刻删除。
如需转载请保留出处:https://51itzy.com/kjqy/71320.html